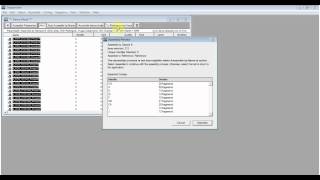
Quickly perform a reference guided alignment using BWA-MEM or GSNAP, and then analyze those alignments for variants using SAMtools Variant Calling. Sequencher can just as easily analyze the entire human genome as it can targeted PCR data. What would analyzing colon cancer data look like using our latest NGS features? It would look a lot like this: Next Gen data is equally easy to analyze with Sequencher thanks to its host of integrated DNA-Seq Tools. Sequencher’s feature list doesn’t end with Sanger data. Instead of combing through each trace peak, you can click one button and have all of the heterozygous bases found and flagged. This feature allows you to set your secondary trace call threshold, what happens to the base call, and automatically marks the heterozygous bases in the contig. Single Nucleotide Polymorphisms are automatically called and clearly displayed in the table, but what about a heterozygote? Sequencher brings in yet another proprietary feature, Calling Secondary Peaks. Below is a screenshot showing an MSH2 analysis in the Variance Table. All four panes are linked: clicking on a base in one will highlight your selection in all four. Using the Review Mode you can simultaneously view the Variance Table, the contig alignment, the corresponding chromatogram traces, and the Translated Variance Table. This table empowers you to quickly and accurately find all of the variants in your data.

Once the alignment is complete, you’ll have access to one of Sequencher’s most powerful features, the Variance Table. Choices range from a simple majority rule to a rigorous forensic standard. Sequencher gives you several options for how the resulting consensus bases are called. You also have the option to adjust the parameters for each algorithm and save them as a template, saving you time on the next project. Each one is tuned for varying levels of sequence quality and genetic variation. Sequencher has three different alignment algorithms for assembling Sanger data. You’ll be amazed at the plethora of tools available, including proprietary algorithms you won’t find anywhere else. The feature list that Sequencher offers for analyzing Sanger data is robust.
CHROMATOGRAM VIEWER SEQUENCHER SOFTWARE
One of the earliest studies linking the Human Mutator Gene Homolog MSH2 to colon cancer featured Sequencher as the software of choice. Sequencher’s powerful algorithms and ease of use have made it a staple for genetic researchers all over the world. The three versions 1.2.1, 2.1.2, and 3.3 are discussed.Researchers have been finding variants and mutations using Sequencher software for a long time. This review explains the general operation of Analysis in terms of viewing and editing a chromatogram, retracking the lanes of a Gel File, and analyzing the final sample data. The sample data are then reextracted from the Gel File and analyzed again. Occasionally the tracking lines within the gel image may need to be adjusted or moved. Ambiguous base calls tend to occur near the end of the sequence and may be either edited or deleted by the user before exporting the data for further comparisons or alignments.


Generally, the products of a sequencing reaction are easily resolved and the Analysis software interprets the correct nucleotide sequence. Individual Sample Files are stored for each of the samples analyzed and include the chromatogram, raw data, and annotations and information regarding the sample and sequence run. This image allows the user to view and edit the tracking lines generated and used by Analysis to collect data points for each sample. The Gel File consists of a computer generated image of the sequencing gel with the fluorescent DNA banding patterns. For the user, there are basically two types of files that can be manipulated to potentially improve the analysis results. After the data are collected from a sequencing run, the Analysis program identifies and tracks the sample lanes of the gel and subsequently normalizes and integrates the raw data into a chromatogram of the final sequence. The ABI sequencer is a laser-based instrument that utilizes fluorescent labels to analyze the products of a sequencing reaction as they migrate through a gel. The ABI Sequencing Analysis application is designed specifically for the analysis of data produced by the ABI DNA Sequencer.
